best-practice variant calling pipeline for automated sequencing analysis; scipy 2013 presentation
Published 11 years ago • 2K plays • Length 6:34Download video MP4
Download video MP3
Similar videos
-
 48:03
48:03
wgs variant calling: variant calling with gatk - part 1 | detailed ngs analysis workflow
-
 4:12
4:12
automated workflow for sequencing data analysis - dhanaprakash jambulingam - la 2022
-
 57:26
57:26
the need for speed: ultra rapid genome sequencing for critical care | dom grand rounds | 24 aug 2022
-
 54:20
54:20
mpg primer: high throughput sequencing and variant calling pipelines (2017)
-
 20:41
20:41
training an unbeatable ai in trackmania
-
 5:01
5:01
how to make a drip irrigation system working model
-
 14:37
14:37
sanger dna sequencing, from then to now.
-
 41:00
41:00
implementing molecular sample tracking in genomic analysis workflows : a case study
-
 14:36
14:36
binf6430grp3pptpresentation
-
 15:03
15:03
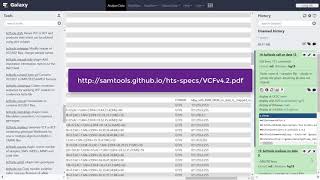
4.4. next generation sequencing - practice session : variant calling
-
 34:13
34:13
small variant calling and annotation
-
 1:35:11
1:35:11
it’s all in the preparation: how to achieve the best hifi data from your samples with pacbio
-
 50:45
50:45
mpg primer: sequence variant calling and data handling (2018)
-
 8:26
8:26
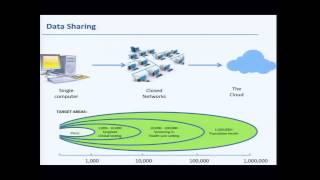
presentation from large-scale genome sequencing and analysis centers' investigators - richard gibbs
-
 7:10
7:10
variant calling tutorial
-
 7:22
7:22
whole genome sequencing (wgs)
-
 56:21
56:21
getting started with whole genome mapping and variant calling on the command line
-
 1:31
1:31
what is sanger sequencing?
-
 27:53
27:53
workflow for comprehensive detection and prioritization of variants with pacbio hifi readsbio
-
 9:12
9:12
13. callset evaluation