how to convert a pdb file into an icm object.
Published 14 years ago • 1.1K plays • Length 1:47Download video MP4
Download video MP3
Similar videos
-
 0:21
0:21
how to save and icm object in pdb format.
-
 38:43
38:43
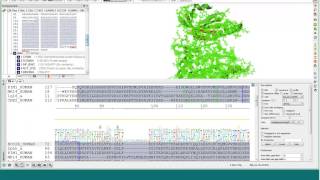
linking protein sequence to 3d structure using the icm alignment editor
-
 2:30
2:30
convert pdb structure into icm object
-
 1:13
1:13
how to extract a 2d sketch of a ligand in complex with a pdb structure.
-
 4:42
4:42
how to re-dock a ligand using molsoft's icm-pro desktop modeling software.
-
 38:20
38:20
application of recyclable thermoplastic composites | piif
-
 4:22
4:22
hiyi milk eco how to connect the printer with milk analyzer
-
 3:09
3:09
mettler toledo ms-ts lab balances - how to setup a connection to an ftp server to export xml files
-
 45:46
45:46
protein loop modeling in icm-pro
-
 1:36
1:36
how to display the ligand binding pocket surface
-
 0:45
0:45
how to search the pdb.
-
 1:00:24
1:00:24
icm-pro: protein structure modeling and analysis
-
 6:25
6:25
convert sdf into pdb file : pymol, openbabel, chimera
-
 44:50
44:50
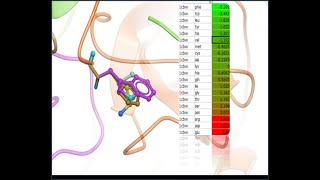
predicting the effect of mutation on protein stability and binding
-
 0:38
0:38
how to save a 2d sketch into a chemical spreadsheet.
-
 2:41
2:41
how to sketch chemicals in the icm molecular editor.
-
 2:38
2:38
how to change protein representation, xstick, ribbon, cpk etc...
-
 43:06
43:06
protac modeling in molsoft's icm-pro software.
-
 6:45
6:45
peptide docking in molsoft's icm-pro
-
 0:51
0:51
how to display energy atomic circles in the icm 3d ligand editor.
-
 52:14
52:14
multi template modeling and editing in molsoft's icm-pro software
-
 2:26
2:26
how to save a docked receptor ligand complex as pdb via the gui and command line.