how to process bottom-up proteomics data with maxquant
Published 5 years ago • 5.1K plays • Length 13:21Download video MP4
Download video MP3
Similar videos
-
 1:01:31
1:01:31
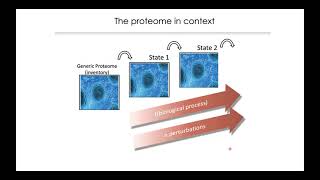
mqss 2021 | translating proteomic data into biological knowledge | ruedi aebersold
-
 24:43
24:43
proteomics focused bioinformatics workshop 2021 - maxquant output and limma results
-
 1:03:11
1:03:11
practical considerations in the design of bottom-up mass spectrometry based proteomics methods
-
 57:27
57:27
lfq-analyst: an interactive platform to analyse & visualise proteomics data processed with maxquant
-
 5:23
5:23
bottom-up proteomics and top-down proteomics
-
 34:16
34:16
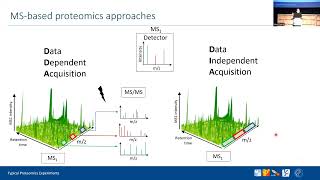
mqss 2022 | typical proteomics experiments | pelagia kyriakidou
-
 48:15
48:15
mqss 2018 | l22: new technology for rapid and comprehensive proteome analysis | joshua coon
-
 36:11
36:11
mqss 2018 | l20: peptide ms/ms spectrum prediction using deep learning | peter cimermancic
-
 39:59
39:59
broade: quantitative methods in proteomics
-
 59:48
59:48
getting started with proteomics
-
 1:58:28
1:58:28
proteomics data analysis
-
 53:36
53:36
mqss 2021 | new mass spectrometer technology for large-scale multi-omic profiling | joshua j. coon
-
 19:25
19:25
peaks studio: protein identification and quantification tutorial
-
 39:23
39:23
mqss 2019 | l9: maxquant output tables | nagarjuna nagaraj
-
 50:38
50:38
mqss 2023|global detection of human variants & isoforms by deep proteome sequencing|pavel sinitcyn
-
 51:06
51:06
mqss 2018 | t3: protein quantification with maxquant | christoph wichmann
-
 31:46
31:46
mqss 2019 | l10: dependent peptides | shivani tiwary
-
 22:42
22:42
mqss 2019 | l16: advanced applications - isobaric labeling | sung-huan yu
-
 17:01
17:01
gtn training - proteomics - introduction to mass spectrometry based proteomics data analysis (old)
-
 51:40
51:40
mqss 2018 | t6: interaction proteomics | jan rudolph
-
 1:02:52
1:02:52
mqss 2019 | t3: how to run maxquant | nagarjuna nagaraj
-
 58:22
58:22
maxquant analysis in galaxy