how to process single cell rnaseq with 2 lines of code from fastq to count matrix 🧬
Published 1 year ago • 1.7K plays • Length 17:06Download video MP4
Download video MP3
Similar videos
-
 5:50
5:50
introduction to single-cell rna-seq and seurat | bioinformatics for beginners
-
 26:09
26:09
a beginner's bioinformatics guide for single-cell rnaseq data analysis
-
 9:36
9:36
processing single-cell rnaseq counts with simpleaf (alevin-fry)
-
 1:50:36
1:50:36
rna-seq i analysis- day 1
-
 13:26
13:26
how to calculate fold change fc, log2fc, pvalue, padj, up and down regulated genes
-
 9:16
9:16
1. what is single cell and why does it matter?
-
 22:20
22:20
rna-seq: introduction and processing fastq files for analysis - pine biotech
-
 7:42
7:42
automated interpretation of single-cell rna-seq data using reference atlases
-
 35:51
35:51
single cell rna-seq | data analysis - dr. raghavendran l. (bioinformatics mentor, pine biotech)
-
 32:14
32:14
how to create pseudo bulk from single cell rnaseq data
-
 20:10
20:10
single cell rna sequencing (scrnaseq): analytical approaches to complex data
-
 6:01
6:01
part2: rna-seq data analysis from fastq files to master regulators
-
 1:18:40
1:18:40
complete single-cell rnaseq analysis walkthrough | advanced introduction
-
 49:41
49:41
qiaseq upx kits for 3’ rnaseq for low input and single cell samples
-
 36:18
36:18
how to analyze single-cell rna-seq data in r | detailed seurat workflow tutorial
-
 24:37
24:37
single cell sequencing - eric chow (ucsf)
-
 17:41
17:41
single cell rna sequencing overview | scrna seq vs bulk seq | chemistry of scrna seq |bio techniques
-
 2:35
2:35
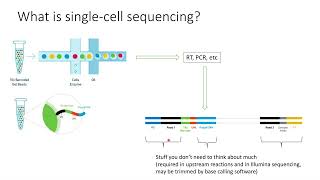
single-cell sequencing explained in 2 minutes
-
 44:56
44:56
introduction to bulk rna-seq
-
 33:26
33:26
single cell gene expression: new insights through the lens of full length mrna isoform resolution
-
 1:57:36
1:57:36
msi live tutorial: rna sequencing analysis 11/05/19