live r coding session - normalizing spatial transcriptomics data for clustering vs deconvolution
Published 1 year ago • 995 plays • Length 34:54Download video MP4
Download video MP3
Similar videos
-
 1:49:47
1:49:47
live r coding session - spatial transcriptomics data analysis with stdeconvolve and spotclean
-
 47:33
47:33
live r coding session - single cell spatial transcriptomics data visualization in base r
-
 6:55
6:55
10x visium spatial transcriptomics data analysis with stdeconvolve in r
-
 20:16
20:16
bioinformatics live coding session - alignment of spatial transcriptomics data using stalign
-
 9:34
9:34
reference-free cell type deconvolution of spatial transcriptomics data with stdeconvolve
-
 1:18:40
1:18:40
complete single-cell rnaseq analysis walkthrough | advanced introduction
-
 28:11
28:11
webinar: retrieving exon and coding region sequences for genes
-
 36:18
36:18
how to analyze single-cell rna-seq data in r | detailed seurat workflow tutorial
-
 26:35
26:35
spatially-resolved transcriptomics analysis with r/bioconductor and beyond
-
 2:28:44
2:28:44
w31: spatial transcriptomics – day 1
-
 17:33
17:33
unsupervised analyses of spatially-resolved transcriptomics data with {nnsvg} and r/bioconductor
-
 9:56
9:56
introduction to spatial sequencing data analysis
-
 47:43
47:43
analysis using r 2023 | 01: exploratory data analysis and introducing clustering
-
 11:39
11:39
data-driven identification of total rna expression genes (tregs)
-
 16:33
16:33
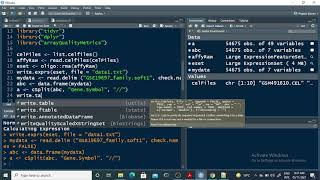
microarray data normalization and annotation - r tutorial
-
 49:56
49:56
visualization of rna sequencing data with pca clustering and heatmaps in rr studio clean
-
 42:51
42:51
ben raphael | alignment, integration, and modeling of spatial transcriptomics data