candidate_genes.m tutorial 02 shortened gene expression table
Published 3 years ago • 40 plays • Length 13:18Download video MP4
Download video MP3
Similar videos
-
 19:46
19:46
deseq2 workflow tutorial | differential gene expression analysis | bioinformatics 101
-
 16:02
16:02
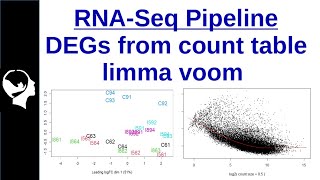
deg isolation using limma voom | a rstudio tutorial
-
 1:00
1:00
differential gene expression (dge) analysis for rna-seq and microarray data using r
-
 26:48
26:48
tutorial: rna-seq workflow with galaxy | no coding involved (step-by-step)
-
 11:21
11:21
introduction to weighted gene co-expression network analysis (wgcna) | bioinformatics 101
-
 48:10
48:10
how to perform gsea - a tutorial on gene set enrichment analysis for rna-seq
-
 28:02
28:02
edger: differential analysis of sequence read count data
-
 1:17:04
1:17:04
webinar #7 – introduction to weighted gene co-expression network analysis
-
 13:26
13:26
how to calculate fold change fc, log2fc, pvalue, padj, up and down regulated genes
-
 35:06
35:06
end-to-end analysis of a gene microarray dataset | bespokeds
-
 9:25
9:25
microarray normalization and differential expression using r
-
 30:36
30:36
differential gene expression analysis in r with deseq2| bioinformatics tutorial for beginners
-
 32:44
32:44
intro to bioinformatics - edger pt 1 - vdb computational biology
-
 9:02
9:02
bioinformatics 101 | how to download rna-seq data from ncbi geo | bioinformatics for beginners
-
 1:19:30
1:19:30
deseq2 workflow tutorial | differential gene expression analysis | rna seq
-
 24:54
24:54
candidate_genes.m tutorial 05 explaining the results: gene expression calculations
-
 41:31
41:31
download data from gdc portal using tcgabiolinks r package